Automated Image Analysis
A scalable solution for reproducible results
· Easily train AI models
· No coding required
· Work with large data sets
· Automate pipelines
arivis and APEER go ZEISS
New names, new look, same outstanding performance
The expanded ‘ZEISS arivis’ product line will include AI and Cloud solutions under new names.
- APEER is now ZEISS arivis Cloud
- The Machine Learning toolkit is now arivis AI The rebranding will happen in phases throughout 2023.
To learn more go to https://www.arivis.com/arivis-news/rebranding-arivis
Faster, reliable results powered by AI
Accelerate your R&D with Automated Image Analysis
Faster, reliable results powered by AI
Why our Customers Choose arivis Cloud
Image processing made easy
Rapid results
Maximum value for money
Support & guidance from our experts
Deep Learning Segmentation
Use deep learning to solve your image analysis task. Take advantage of the full power of deep learning without any coding knowledge. Our user friendly UI guides you through the required steps to train your own model using your own data.


1. Annotate
As a subject matter expert, define the ground truth by painting a few pixels in your training images.

2. Train
To start the deep learning training, just click the Train button.

3. Segment
Use the trained model to segment new images or download the model for offline image analysis.
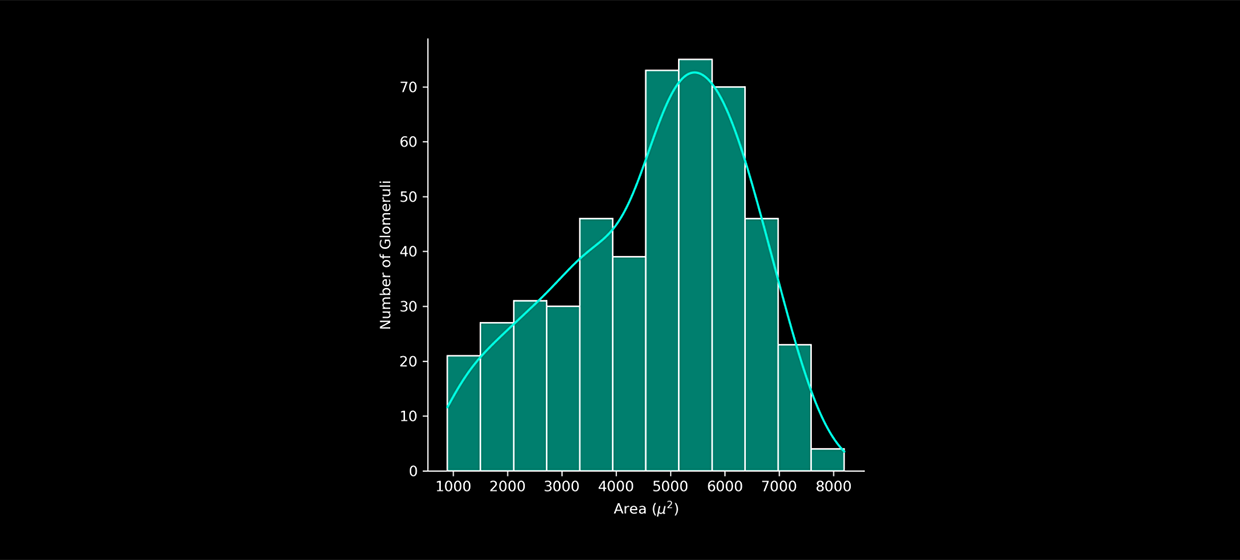
4. Insights
Use the reported post-segmentation metrics to gain insights from your images.
Automated Pipelines
Fully automate routine image analysis using AI-trained models to boost your productivity and ensure reproducible results. Automated pipelines can be customized for your specific requirements using the ZEISS digital ecosystem.






























